Critical Spectrum Calculations
Return to Diffusion Coefficients and Critical Spectrum Search documentation.
import matplotlib.pyplot as plt
from matplotlib import rcParams
# Default values
FONT_SIZE = 16 # font size for plotting purposes
# rcParams['figure.dpi'] = 300
plt.rcParams['figure.figsize'] = [6, 4] # Set default figure size
import numpy as np
import serpentTools
from scipy.linalg import solve
from plotter import Plot1d
from critical_spectrum import SolveB1, SolveCM, CriticalSpectrum, Condense2gr
The \(B_1\) Critical spectrum
Read results from Serpent
resFile70gr = "./serpent/fa2D_70gr_inf_res.m"
resFile2gr = "./serpent/fa2D_2gr_B1_res.m"
Two-group results [reference point]
res2gr = serpentTools.read(resFile2gr)
univ = res2gr.getUniv('0', timeDays=0)
b1Transp_2gr = univ.b1Exp['b1Transpxs']
b1Rabsxs_2gr = univ.b1Exp['b1Rabsxs']
infTransp_2gr = univ.infExp['infTranspxs']
infRabsxs_2gr = univ.infExp['infRabsxs']
infDiff_2gr = 1/(3*infTransp_2gr)
b1Diff_2gr = 1/(3*b1Transp_2gr)
SERPENT Serpent 2.2.1 found in ./serpent/fa2D_2gr_B1_res.m, but version 2.1.31 is defined in settings
Attempting to read anyway. Please report strange behaviors/failures to developers.
70-group results
# Read the results using the serpentTools package
res70gr = serpentTools.read(resFile70gr)
SERPENT Serpent 2.2.1 found in ./serpent/fa2D_70gr_inf_res.m, but version 2.1.31 is defined in settings
Attempting to read anyway. Please report strange behaviors/failures to developers.
ng = 70 # number of energy groups
univ0 = res70gr.getUniv('0', timeDays=0)
flx = univ0.infExp['infFlx']
sigT = univ0.infExp['infTot']
rabsxs = univ0.infExp['infRabsxs']
nusigF = univ0.infExp['infNsf']
chi = univ0.infExp['infChit']
SP1 = univ0.infExp['infSp1']
SP0 = univ0.infExp['infSp0']
cmmTransp = univ0.gc['cmmTranspxs'] # represents in-scatter
infTransp = univ0.infExp['infTranspxs'] # out-scatter
energy = univ0.groups * 1E+06
kinf = res70gr.resdata['absKeff'][0] # k-inf
To create the scattering matrix with the following structure: 1->1, 2->1 3->1 … 1->2, 2->2 3->2 … 1->3, 2->3 3->3 … …
SP1=SP1.reshape((ng,ng)).transpose()
SP0=SP0.reshape((ng,ng)).transpose()
Solving \(B_1\) equations with a known \(\alpha_g\) and \(B^2\)
From the second \(B_1\) equation a relation between \(J\) and \(\phi\) is established.
\(\pm iB \vec{J} = B^2 \mathbf{D} \vec{\phi}\)
Components of the inverse of D are:
\(D_{gg'}^{-1}=3\alpha_g\Sigma_{t,g}\delta_{gg'}-3\Sigma_{s1,g'g}\)
\(\vec{\phi} = \mathbf{A}^{-1}~\vec{\chi}\)
where, the components of matrix \(A\) are:
\(A_{gg'}=\Sigma_{t,g}\delta_{gg'}-\Sigma_{s0,g'g}+B^2D_{gg'}\)
Solve \(B_1\)
Methodology to iterate on \(B_1\)
\(B^2\)=0 is used to obtain the infinite-medium \(k_{\infty}\) and its associated spectrum. Serpent provide \(k_{\infty}\).
A perturbed value \(B^2=0.0001\) is used to solve for the flux and current.
The coefficient \(\frac{k_{\infty}}{M^2}\) is evaluated only once.
Energy condensation
Execute the entire sequence
crit_flxB1, crit_iJB1, B2B1, iB1 = CriticalSpectrum(ng, SP0, SP1, sigT, cmmTransp, chi, nusigF, kinf, P1=False, CM=False)
crit_flxP1, crit_iJP1, B2P1, iP1 = CriticalSpectrum(ng, SP0, SP1, sigT, cmmTransp, chi, nusigF, kinf, P1=True, CM=False)
crit_flxCM, B2CM, iCM= CriticalSpectrum(ng, SP0, SP1, sigT, cmmTransp, chi, nusigF, kinf, P1=False, CM=True)
print(f"The critical B^2 values are B1:{B2B1:.6f}, P1:{B2P1:.6f}, CM:{B2CM:.6f}.")
print(f"k_inf = 1 +/- 1e-6 was achieved in B1:{iB1} iterations, P1:{iP1} iterations, and CM:{iCM} iterations.")
The critical B^2 values are B1:0.000255, P1:0.000254, CM:0.000246.
k_inf = 1 +/- 1e-6 was achieved in B1:4 iterations, P1:3 iterations, and CM:3 iterations.
normFlx = flx / flx.sum()
normFlxCM = crit_flxCM / crit_flxCM.sum()
normFlxB1 = crit_flxB1 / crit_flxB1.sum()
normFlxP1 = crit_flxP1 / crit_flxP1.sum()
flx_diff = 100*(1-normFlx/normFlxB1)
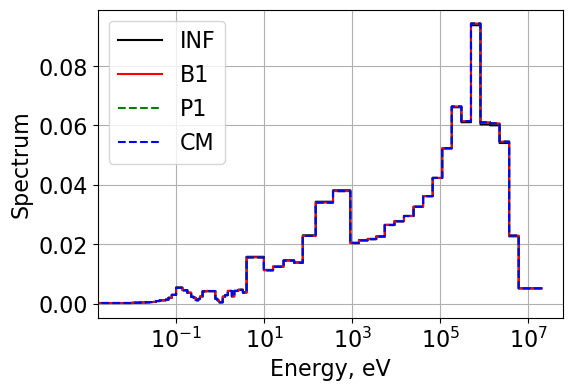
Compare the flux and current spectrum
plt.figure()
plt.grid(visible=True)
Plot1d(energy, normFlx, xlabel="Energy, eV", ylabel='',
fontsize=16, marker="-k", markerfill=False, markersize=4)
Plot1d(energy, normFlxB1, xlabel="Energy, eV", ylabel='Spectrum',
fontsize=16, marker="-r", markerfill=False, markersize=6)
Plot1d(energy, normFlxP1, xlabel="Energy, eV", ylabel='Spectrum',
fontsize=16, marker="--g", markerfill=False, markersize=6)
Plot1d(energy, normFlxCM, xlabel="Energy, eV", ylabel='Spectrum',
fontsize=16, marker="--b", markerfill=False, markersize=4)
plt.legend(['INF', 'B1',"P1",'CM'])
<matplotlib.legend.Legend at 0x215bd737190>
Diffusion coefficients
# Use eq. 8.21 to obtain the diffusion coefficient
DgB1 = (crit_iJB1) / (crit_flxB1 * B2B1**0.5)
DgP1 = (crit_iJP1) / (crit_flxP1 * B2P1**0.5)
# Note that the diffusion coefficients used in the CM method are calculated using cmmTransport cross sections from Serpent output
Condense into 2-group
pred_b1Rabsxs_2gr = Condense2gr(rabsxs, crit_flxB1, energy, cutoffE=0.625)
pred_b1Diff_2gr = Condense2gr(DgB1, crit_flxB1, energy, cutoffE=0.625)
pred_p1Rabsxs_2gr = Condense2gr(rabsxs, crit_flxP1, energy, cutoffE=0.625)
pred_p1Diff_2gr = Condense2gr(DgP1, crit_flxP1, energy, cutoffE=0.625)
pred_cmRabsxs_2gr = Condense2gr(rabsxs, crit_flxCM, energy, cutoffE=0.625)
pred_cmDiff_2gr = Condense2gr(1/(3*cmmTransp), crit_flxCM, energy, cutoffE=0.625)
print(f"B1 (Serpent, Reference) results, Dg: {b1Diff_2gr}, Rabsxs: {b1Rabsxs_2gr}.")
print(f"B1 2-group results, Dg: {pred_b1Diff_2gr}, Rabsxs: {pred_b1Rabsxs_2gr}.")
print(f"P1 2-group results, Dg: {pred_p1Diff_2gr}, Rabsxs: {pred_p1Rabsxs_2gr}.")
print(f"CM 2-group results, Dg: {pred_cmDiff_2gr}, Rabsxs: {pred_cmRabsxs_2gr}.")
B1 (Serpent, Reference) results, Dg: [1.69958768 0.88299987], Rabsxs: [0.00775962 0.0676255 ].
B1 2-group results, Dg: [1.69905265 0.88293977], Rabsxs: [0.00776182 0.06764274].
P1 2-group results, Dg: [1.70307681 0.88330774], Rabsxs: [0.00776193 0.06764278].
CM 2-group results, Dg: [1.76303046 0.88447132], Rabsxs: [0.00776121 0.0676434 ].