AFEN Method
Return to Analytic Function Expansion Nodal Method documentation.
from IPython.display import Image
import matplotlib.pyplot as plt
from matplotlib import rcParams
import numpy as np
import time
# Default values
FONT_SIZE = 16 # font size for plotting purposes
# rcParams['figure.dpi'] = 300
plt.rcParams['figure.figsize'] = [6, 4] # Set default figure size
from analytic_nodal_expansion import AFEN2D, meshPlot, GetSerpentRes
from NEM import CartesianNem1D, Plot1d
import serpentTools
Flux and pin-power reconstruction
Colorset 2-dim test case
imgFile = './serpent/SMR/SMR_Ref_2D_2g_geom1.png'
resFile = './serpent/SMR/SMR_Ref_2D_2g_res.m'
detFile = './serpent/SMR/SMR_Ref_2D_2g_det0.m'
Image(imgFile, width=400, height=300)
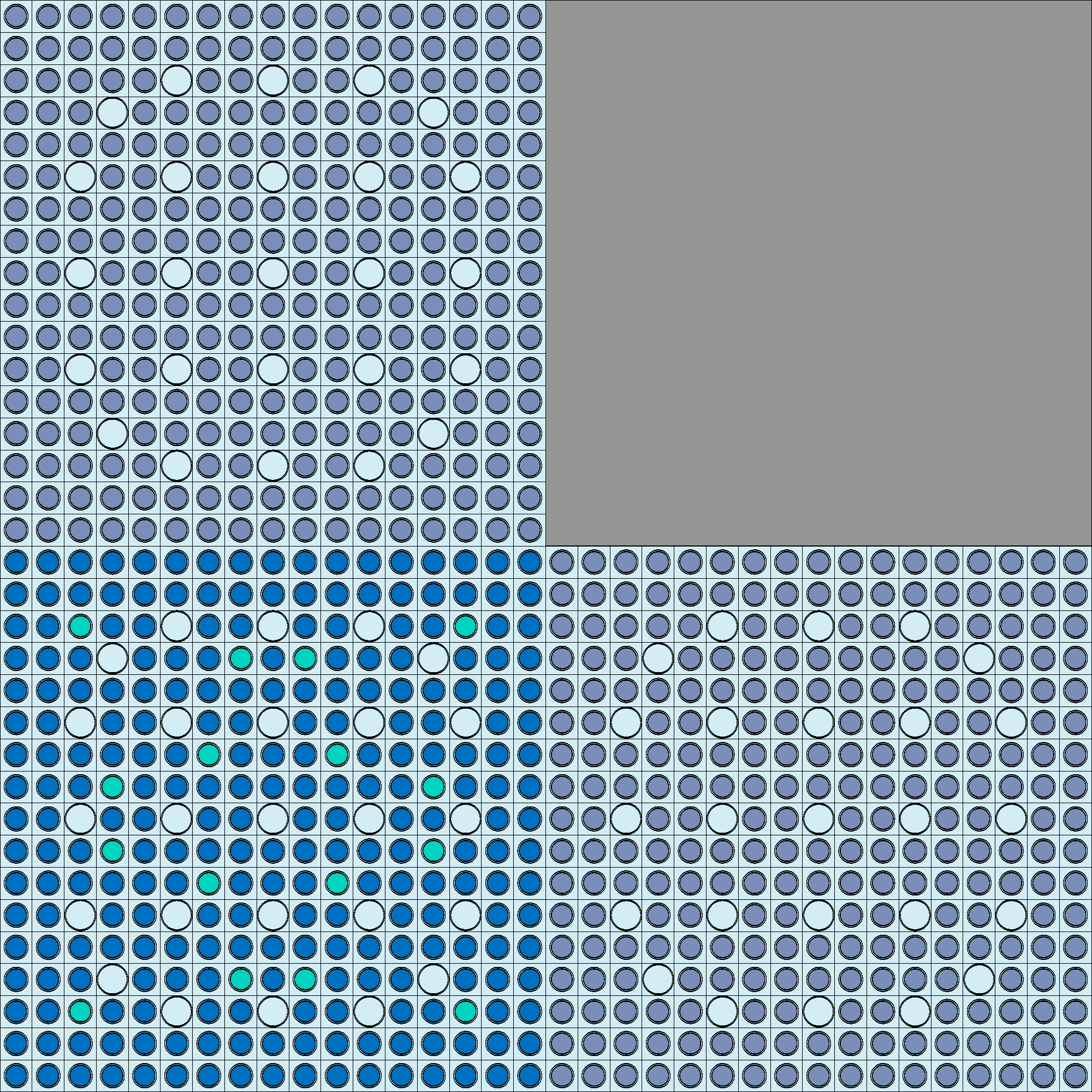

Universes within the colorset: - Upper left F12 - 2.6% \(UO_2\) - Upper right Ref - stainless steel (with water for PWR) - Bottom left F2 - 4.55% \(UO_2\) with 8% \(Gd_2\) \(O_3\) - Bottom right F11 - 2.6% \(UO_2\)
Read detector and results data from Serpent
det = serpentTools.read(detFile)
xsF2, bcF2 = GetSerpentRes(resFile, 'F2', 0)
xsF11, bcF11 = GetSerpentRes(resFile, 'F11', 0)
xsF12, bcF12 = GetSerpentRes(resFile, 'F12', 0)
xsRef, bcRef = GetSerpentRes(resFile, 'Ref', 0)
SERPENT Serpent 2.2.1 found in ./serpent/SMR/SMR_Ref_2D_2g_res.m, but version 2.1.31 is defined in settings
Attempting to read anyway. Please report strange behaviors/failures to developers.
SERPENT Serpent 2.2.1 found in ./serpent/SMR/SMR_Ref_2D_2g_res.m, but version 2.1.31 is defined in settings
Attempting to read anyway. Please report strange behaviors/failures to developers.
SERPENT Serpent 2.2.1 found in ./serpent/SMR/SMR_Ref_2D_2g_res.m, but version 2.1.31 is defined in settings
Attempting to read anyway. Please report strange behaviors/failures to developers.
SERPENT Serpent 2.2.1 found in ./serpent/SMR/SMR_Ref_2D_2g_res.m, but version 2.1.31 is defined in settings
Attempting to read anyway. Please report strange behaviors/failures to developers.
Store cross sections and bc in dicts
bc = {'F2': bcF2, 'F11': bcF11, 'F12': bcF12, 'Ref': bcRef,}
xs = {'F2': xsF2, 'F11': xsF11, 'F12': xsF12, 'Ref': xsRef,}
dx, dy = 21.42, 21.42 # cm - assembly length
npins = 17
Manipulate the data provided by Serpent (BC)
Serpent already provodes the het flux values for all the surfaces and corners for each universe.
bcflux = {}
for univ in bc: # defining BC for each universe
bcflux[univ] = {}
# store the surface and corner fluxes
bcflux[univ]['av'] = bc[univ]['flux']
bcflux[univ]['w'] = bc[univ]['wFlux']
bcflux[univ]['e'] = bc[univ]['eFlux']
bcflux[univ]['s'] = bc[univ]['sFlux']
bcflux[univ]['n'] = bc[univ]['nFlux']
# bcflux[univ]['nw'] = bc[univ]['nwFlux']
# bcflux[univ]['ne'] = bc[univ]['neFlux']
# bcflux[univ]['sw'] = bc[univ]['swFlux']
# bcflux[univ]['se'] = bc[univ]['seFlux']
bcflux[univ]['nw'] = bcflux[univ]['n']+bcflux[univ]['w']-bcflux[univ]['av']
bcflux[univ]['ne'] = bcflux[univ]['n']+bcflux[univ]['e']-bcflux[univ]['av']
bcflux[univ]['sw'] = bcflux[univ]['s']+bcflux[univ]['w']-bcflux[univ]['av']
bcflux[univ]['se'] = bcflux[univ]['s']+bcflux[univ]['e']-bcflux[univ]['av']
Manually average the corner fluxes using adjacent fuel assemblies
bcflux['F12']['ne'] = (bcflux['F12']['ne'] + bcflux['Ref']['nw'])/2
bcflux['Ref']['nw'] = bcflux['F12']['ne']
bcflux['F12']['sw'] = (bcflux['F12']['sw'] + bcflux['F2']['nw'])/2
bcflux['F2']['nw'] = bcflux['F12']['sw']
bcflux['F2']['se'] = (bcflux['F2']['se'] + bcflux['F11']['sw'])/2
bcflux['F11']['sw'] = bcflux['F2']['se']
bcflux['F11']['ne'] = (bcflux['F11']['ne'] + bcflux['Ref']['se'])/2
bcflux['Ref']['se'] = bcflux['F11']['ne']
bcflux['F12']['se'] = (bcflux['F12']['se'] + bcflux['Ref']['sw']+
bcflux['F2']['ne'] + bcflux['F11']['nw'])/4
bcflux['Ref']['sw'] = bcflux['F12']['se']
bcflux['F2']['ne'] = bcflux['F12']['se']
bcflux['F11']['nw'] = bcflux['F12']['se']
Reconstruct the homogeneous flux
univId = 'F2' # user needs to choose
timeStart = time.perf_counter()
univres = AFEN2D(xs[univId], bcflux[univId], dx, symbolic=False)
univres.ReconstructFlux() # a built-in method to obtain the coeffs
timeEnd = time.perf_counter()
print('Reconstructed flux for {} calculated in {} seconds'.format(univId,timeEnd-timeStart))
Reconstructed flux for F2 calculated in 0.001218500008690171 seconds
yvals = np.linspace(-dx/2, +dx/2, npins+1)
yvals = 0.5*(yvals[1:]+yvals[0:-1])
timeStart1 = time.perf_counter()
univres.GetFlux2D(yvals, yvals)
timeEnd1 = time.perf_counter()
print('2D neutron flux for {} calculated in {} seconds'.format(univId,timeEnd1-timeStart1))
2D neutron flux for F2 calculated in 0.005599600001005456 seconds
Reference flux from Serpent
if univId == 'F2':
fastHetFlux = det.detectors['flux_fast'].tallies[0:npins, 0:npins]
thermalHetFlux = det.detectors['flux_thermal'].tallies[0:npins, 0:npins]
fastPower = det.detectors['power_fast'].tallies[0:npins, 0:npins]
thermalPower = det.detectors['power_thermal'].tallies[0:npins, 0:npins]
elif univId == 'F11':
fastHetFlux = det.detectors['flux_fast'].tallies[0:npins, npins:]
thermalHetFlux = det.detectors['flux_thermal'].tallies[0:npins, npins:]
fastPower = det.detectors['power_fast'].tallies[0:npins, npins:]
thermalPower = det.detectors['power_thermal'].tallies[0:npins, npins:]
elif univId == 'F12':
fastHetFlux = det.detectors['flux_fast'].tallies[npins:, 0:npins]
thermalHetFlux = det.detectors['flux_thermal'].tallies[npins:, 0:npins]
fastPower = det.detectors['power_fast'].tallies[npins:, 0:npins]
thermalPower = det.detectors['power_thermal'].tallies[npins:, 0:npins]
elif univId == 'Ref':
fastHetFlux = det.detectors['flux_fast'].tallies[npins:, npins:]
thermalHetFlux = det.detectors['flux_thermal'].tallies[npins:, npins:]
fastPower = det.detectors['power_fast'].tallies[npins:, npins:]
thermalPower = det.detectors['power_thermal'].tallies[npins:, npins:]
Plot results
Fast flux distribution
Thermal flux distribution
Fast Flux 2-dim
meshPlot(fastHetFlux, 17, univId, 'flux')
meshPlot(univres.flux2d[0], 17, univId, 'flux')
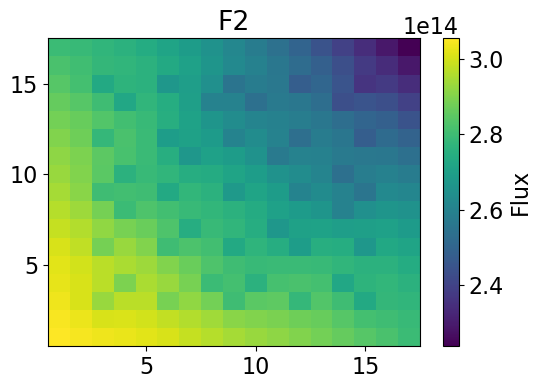
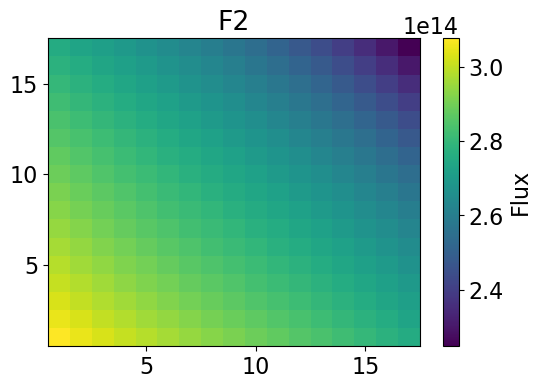
Thermal Flux 2-dim
meshPlot(thermalHetFlux, 17, univId, 'flux')
meshPlot(univres.flux2d[1], 17, univId, 'flux')
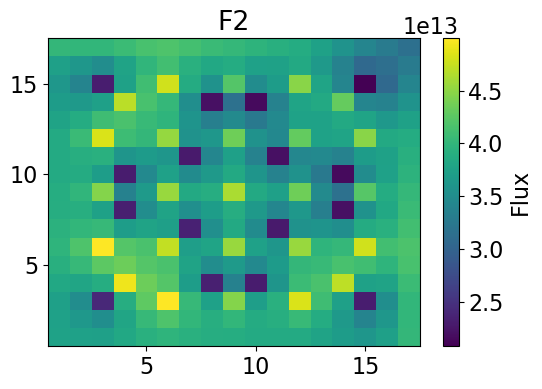
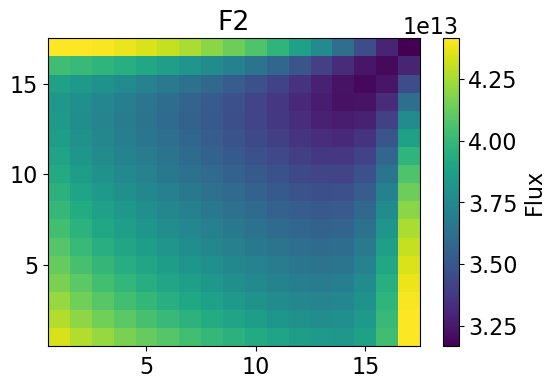
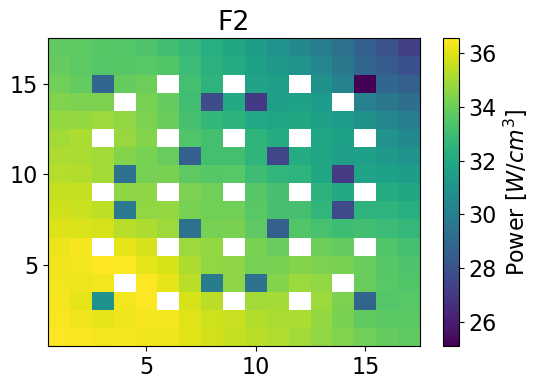
Fast Power 2-dim
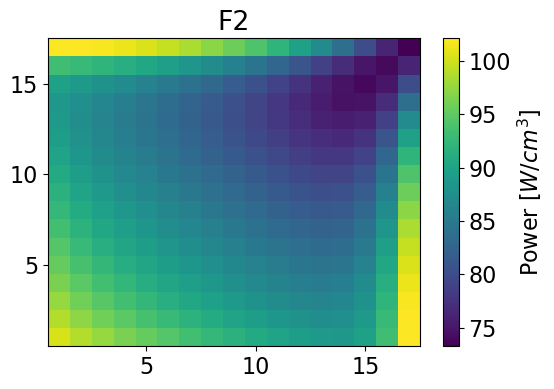
meshPlot(fastPower, 17, univId, 'power')
fastPowerHom = univres.HomogeneousPower(univres.flux2d, npins)[0]
meshPlot(fastPowerHom, 17, univId, 'power')
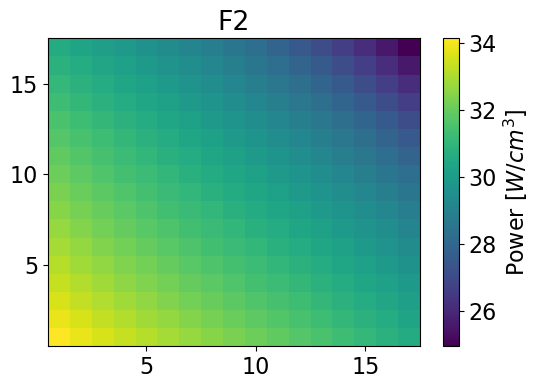
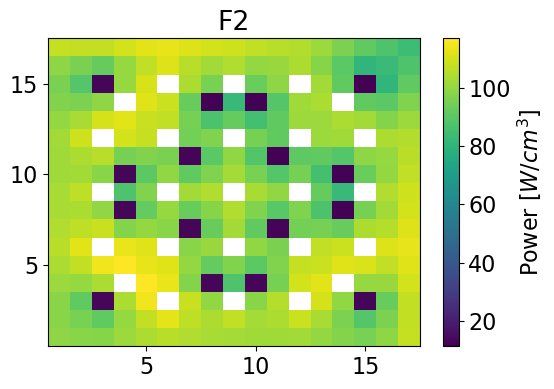
Thermal Power 2-dim
meshPlot(thermalPower, 17, univId, 'power')
thermalPowerHom = univres.HomogeneousPower(univres.flux2d, npins)[1]
meshPlot(thermalPowerHom, 17, univId, 'power')
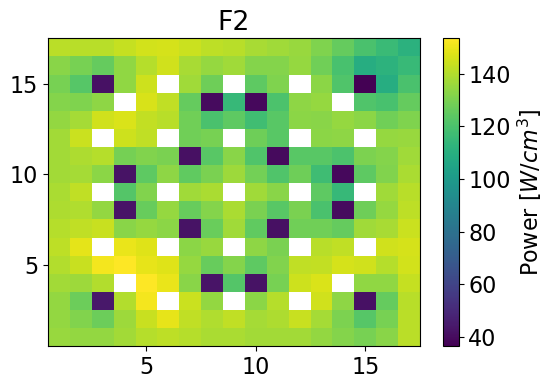
Total Power Plots
serpentPower = fastPower + thermalPower
totalPower = univres.HomogeneousPower(univres.flux2d, npins, total=True)
meshPlot(serpentPower, 17, univId, 'power')
meshPlot(totalPower, 17, univId, 'power')
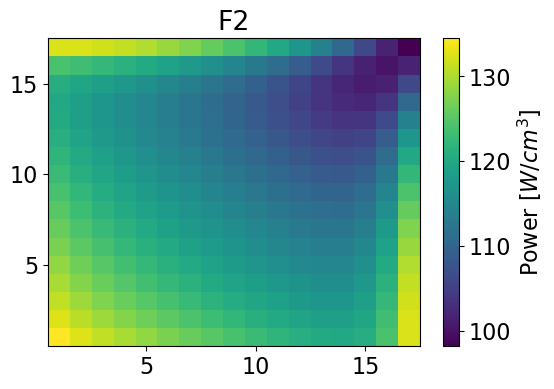
Error for homogenized power distributions
# mask = fastPower != 0
# fastPower[fastPower == 0] = np.nan
# relativeError = np.abs(fastPower - fastPowerHom) / fastPower
# # Mask invalid entries with -inf so they’re ignored in max
# maskedError = np.where(mask, relativeError, -np.inf)
# maxRelativeError = np.max(maskedError) * 100
# avRelativeError = np.mean(relativeError[~np.isnan(relativeError)]) * 100
# maxIdx = np.unravel_index(np.argmax(maskedError), maskedError.shape)
# row, col = int(maxIdx[0]), int(maxIdx[1])
# print(f"Max relative error in fast power: {maxRelativeError:.3f}% at index ({row}, {col}) with an average relative error of: {avRelativeError:.3f}%")
# mask = thermalPower != 0
# thermalPower[thermalPower == 0] = np.nan
# relativeError = np.abs(thermalPower - thermalPowerHom) / thermalPower
# # Mask invalid entries with -inf so they’re ignored in max
# maskedError = np.where(mask, relativeError, -np.inf)
# maxRelativeError = np.max(maskedError) * 100
# avRelativeError = np.mean(relativeError[~np.isnan(relativeError)]) * 100
# maxIdx = np.unravel_index(np.argmax(maskedError), maskedError.shape)
# row, col = int(maxIdx[0]), int(maxIdx[1])
# print(f"Max relative error in thermal power: {maxRelativeError:.3f}% at index ({row}, {col}) with an average relative error of: {avRelativeError:.3f}%")
Plot x-dependent y-averaged results for AFEN and NEM
thermalPowerHomX2 = thermalPowerHom.mean(axis=0)
# solve for second region using AFEN Method
univId = 'F11'
tally = det.detectors['flux_fast'].x[:, 1]
fastHetFlux = det.detectors['flux_fast'].tallies[npins:, :]
thermalHetFlux = det.detectors['flux_thermal'].tallies[npins:, :]
fastPower = det.detectors['power_fast'].tallies[npins:, :]
thermalPower = det.detectors['power_thermal'].tallies[0:npins]
univres = AFEN2D(xs[univId], bcflux[univId], dx, symbolic=False)
univres.ReconstructFlux() # a built-in method to obtain the coeffs
univres.GetFlux2D(yvals, yvals)
thermalPowerHomX11 = univres.HomogeneousPower(univres.flux2d, npins)[1].mean(axis=0)
# build out NEM solutions for two regions
fastHetFluxNEM = det.detectors['flux_fast'].tallies[npins:].mean(axis=0)
thermalHetFluxNEM = det.detectors['flux_thermal'].tallies[npins:].mean(axis=0)
universesNEM = ['F2', 'F11']
xs1, bc1 = GetSerpentRes(resFile, universesNEM[0], timeDays=0)
xs2, bc2 = GetSerpentRes(resFile, universesNEM[1], timeDays=0)
# Left assembly solution
trLeakage1 = {}
trLeakage1['eL'] = bc2['nJnet'] - bc2['sJnet']
trLeakage1['eD'] = xs2['diff']
trLeakage1['edx'] = dx
trLeakage1['wL'] = bc1['nJnet'] - bc1['sJnet']
trLeakage1['wD'] = xs1['diff']
trLeakage1['wdx'] = dx
nem1 = CartesianNem1D(dx, dy, xs1, bc1, trLeakage1, symbolic=False)
nem1.TransverseLeakageCoef('x') # obtain the coefficients of the TL
nem1.GetExpansionCoeffs('x', 'diff')
xvals = np.linspace(-dx/2, +dx/2, npins+1)
xvals = 0.5*(xvals[1:]+xvals[0:-1])
flux1 = nem1.GetHomogFlux(xvals)
# Right assembly solution
trLeakage2 = {}
trLeakage2['wL'] = bc1['nJnet'] - bc1['sJnet']
trLeakage2['wD'] = xs1['diff']
trLeakage2['wdx'] = dx
trLeakage2['eL'] = bc2['nJnet'] - bc2['sJnet']
trLeakage2['eD'] = xs2['diff']
trLeakage2['edx'] = dx
nem2 = CartesianNem1D(dx, dy, xs2, bc2, trLeakage2, symbolic=False)
nem2.TransverseLeakageCoef('x') # obtain the coefficients of the TL
nem2.GetExpansionCoeffs('x', 'diff')
flux2 = nem2.GetHomogFlux(xvals)
SERPENT Serpent 2.2.1 found in ./serpent/SMR/SMR_Ref_2D_2g_res.m, but version 2.1.31 is defined in settings
Attempting to read anyway. Please report strange behaviors/failures to developers.
SERPENT Serpent 2.2.1 found in ./serpent/SMR/SMR_Ref_2D_2g_res.m, but version 2.1.31 is defined in settings
Attempting to read anyway. Please report strange behaviors/failures to developers.
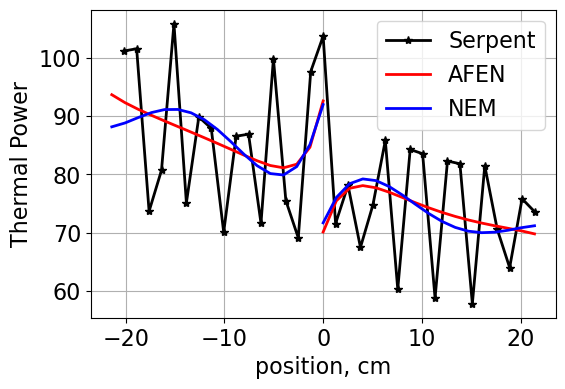
# Plot all methods for comparison
xvals = np.linspace(-dx/2, +dx/2, npins)
plt.figure()
Plot1d(tally, thermalPower.mean(axis=0), xlabel="position, cm",
ylabel='',
fontsize=16, marker="-*k", markersize=6, linewidth=2, label="Serpent")
Plot1d(xvals-10.71, thermalPowerHomX2, xlabel="position, cm",
ylabel='Fast Flux',
fontsize=16, marker="-r", markersize=6, linewidth=2, label="AFEN")
Plot1d(xvals+10.71, thermalPowerHomX11, xlabel="position, cm",
ylabel='Fast Flux',
fontsize=16, marker="-r", markersize=6, linewidth=2, label="_nolegend_")
Plot1d(xvals-10.71, xs1['kappa'][1]*1.60218e-13*xs1['fiss'][1]*flux1[1,:], xlabel="position, cm",
ylabel=None,
fontsize=16, marker="-b", markersize=6, linewidth=2, label='NEM')
Plot1d(xvals+10.71, xs2['kappa'][1]*1.60218e-13*xs2['fiss'][1]*flux2[1,:], xlabel="position, cm",
ylabel='Thermal Power',
fontsize=16, marker="-b", markersize=6, linewidth=2, label="_nolegend_")
plt.legend()
plt.grid(visible=True)